 Despite
what you may think, this isn't a personal comment! Rather it is to draw
your attention to a theory proposed 50 years ago, by Francis Crick in a
publication in the Journal of Molecular Biology in 1966 entitled: Codon--anticodon pairing: the wobble hypothesis. The link takes you to a review
written to celebrate 40 years of wobble; sadly the original paper is
harder to find online. So I will try and explain why I believe it is one
of the most elegant pieces of scientific thinking, from one of the
founding fathers of modern Molecular Biology. It is also timely, since
the 50th Anniversary coincides with the planned opening later this year,
of the Crick Institute close to London's three, great North facing Railway Stations: a metaphor for the triplet nature of the genetic code perhaps?
Despite
what you may think, this isn't a personal comment! Rather it is to draw
your attention to a theory proposed 50 years ago, by Francis Crick in a
publication in the Journal of Molecular Biology in 1966 entitled: Codon--anticodon pairing: the wobble hypothesis. The link takes you to a review
written to celebrate 40 years of wobble; sadly the original paper is
harder to find online. So I will try and explain why I believe it is one
of the most elegant pieces of scientific thinking, from one of the
founding fathers of modern Molecular Biology. It is also timely, since
the 50th Anniversary coincides with the planned opening later this year,
of the Crick Institute close to London's three, great North facing Railway Stations: a metaphor for the triplet nature of the genetic code perhaps? By 1966, Francis Crick had solved the structure of DNA (with the help of Jim
Watson and colleagues including Rosalind Franklin, Maurice Wilkins and
Raymond Gosling) as well as correctly predicting the existence of
transfer RNA and contributing to the elucidation of the genetic code
with his colleague of the time, Sydney Brenner. All of which made Crick
perfectly placed to propose a molecular explanation for the apparent
mismatch between the number of unique amino acids found in proteins (20)
and the number of triplet codons found in mRNA (64, including 3
nonsense codons that instruct the ribosome to stop protein synthesis).
It is also timely, since the ribosome has been on the one hand the
target of many prescription antibiotics, and a source of interest since
its RNAs and proteins, when mutated in specific ways, can shrug off the
challenge of such antibiotics. You can read more about this important
issue in my recent post here.
By 1966, Francis Crick had solved the structure of DNA (with the help of Jim
Watson and colleagues including Rosalind Franklin, Maurice Wilkins and
Raymond Gosling) as well as correctly predicting the existence of
transfer RNA and contributing to the elucidation of the genetic code
with his colleague of the time, Sydney Brenner. All of which made Crick
perfectly placed to propose a molecular explanation for the apparent
mismatch between the number of unique amino acids found in proteins (20)
and the number of triplet codons found in mRNA (64, including 3
nonsense codons that instruct the ribosome to stop protein synthesis).
It is also timely, since the ribosome has been on the one hand the
target of many prescription antibiotics, and a source of interest since
its RNAs and proteins, when mutated in specific ways, can shrug off the
challenge of such antibiotics. You can read more about this important
issue in my recent post here.In 1966 there was a consensus view that Crick's "central dogma" of Molecular Biology: DNA makes RNA makes Protein, was correct. It would take the arrival of the AIDS virus over ten years later to begin to add some caveats, but that won't concern us here. It was also known that there was a mismatch between the number of proteogenic amino acids (not a word I like, but it serves to define the standard set of twenty amino acids that are incorporated into proteins during ribosomal translation, and excludes some of the "fancy" amino acids that are particularly common in plants). Crick explains the need for a solution to this paradox, rather concisely and elegantly in the opening abstract to his 1966 paper in the Journal of Molecular Biology:
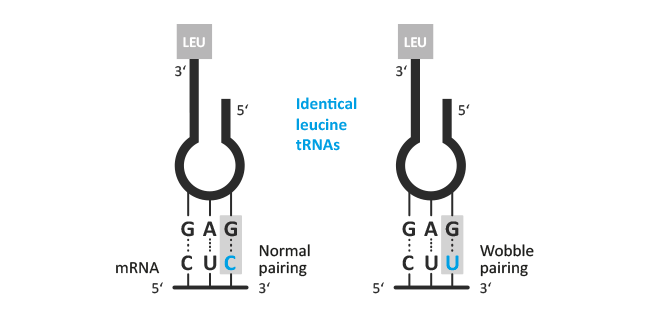
"It is suggested that while the standard base pairs may be used rather strictly
in the first two positions of the triplet, there may be some wobble in the pairing
of the third base. This hypothesis is explored systematically, and it is shown
that such a wobble could explain the general nature of the degeneracy of the
genetic code."
The triplet nature of the code is simple: every amino acid in a protein chain is coded for by three nucleotides present in messenger RNA (mRNA): simple now, but a major challenge in the early 1960s! The coding section of mRNA is decoded by 3 complementary nucleotides in the anticodon triplet that are located at the "tip" of the transfer RNA (tRNA, originally called soluble RNA, or sRNA) molecule. Each codon has an anticodon and each anticodon is physically linked to a specific amino acid. The precise and invariable pairing of anticodon and amino acid is in itself a fascinating aspect of Biological Fidelity and formed the basis of the early career of one of the UK's most distinguished enzymologists, Professor Sir Alan Fersht, now Master of Gonville and Caius College Cambridge. His landmark book contains an extensive discussion of the enzymes that catalyse the high fidelity attachment of an amino acid to a tRNA: the aminoacyl-transfer RNA synthetases.
Occam's Razor would lead us to suppose that there are 20 tRNAs, one per amino acid. Maybe one more for a start codon and another for a stop codon would add two more, but the finding that there are in fact 64 codons was something of a conundrum: and while experimental Molecular Biologists began the search for new tRNAs, Crick applied his (not inconsiderable!) intellect to the problem in a different way. To attempt to understand how the mind of a genius works is a dangerous thing, but we can see in his paper that he draws on a deep understanding of the principles of physical chemistry: bonding, geometry and polarity, for example. These are all combined with a clear appreciation of stereo-chemistry and symmetry; all of which undoubtedly came in handy when he teamed up with Jim Watson in the early 1950s! Before we get to arguments, I can update you and tell you that there are in fact (typically) 40 tRNAs in a cell. Moreover, the start codon is generally (but there are exceptions) the unique codon that specifies the sulphur-containing, hydrophobic amino acid methionine. There are three stop codons, attractively called opal, amber and ochre by bacterial geneticists! So we have 61 proteogenic codons and 40 tRNAs: thus providing a contemporary context to Crick's hypothesis.
No comments:
Post a Comment