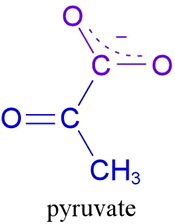
 My choice for September reflects my growing interest in the molecular evolution of enzymes in "simple" organisms like E.coli. Pyruvate Dehydrogenase (PDH for short), catalyses the inter-facial reaction, sometimes called the "linking" reaction, between glycolysis (the conversion of sugars such as glucose to pyruvate) and the ubiquitous Krebs Cycle, in which pyruvate is first converted to acetyl coenzyme A, in preparation for entry into a series of cyclic intercoversions that yield NADH, FADH and either GTP or ATP. You can read about NAD(H) in my previous Molecule of the Month and the early experimental challenges of metabolism here. The conversion of pyruvate (top left), a simple carboxylic acid, to acetyl coenzyme A in most organisms requires a triptych of enzymes: E1 catalyses the oxidative decarboxylation of pyruvate with the help of the cofactor thiamine pyrophosphate (TPP). The other name for TPP, is vitamin B1: an essential nutrient. CO2 is also released in the first step of the PDH reaction(s).
My choice for September reflects my growing interest in the molecular evolution of enzymes in "simple" organisms like E.coli. Pyruvate Dehydrogenase (PDH for short), catalyses the inter-facial reaction, sometimes called the "linking" reaction, between glycolysis (the conversion of sugars such as glucose to pyruvate) and the ubiquitous Krebs Cycle, in which pyruvate is first converted to acetyl coenzyme A, in preparation for entry into a series of cyclic intercoversions that yield NADH, FADH and either GTP or ATP. You can read about NAD(H) in my previous Molecule of the Month and the early experimental challenges of metabolism here. The conversion of pyruvate (top left), a simple carboxylic acid, to acetyl coenzyme A in most organisms requires a triptych of enzymes: E1 catalyses the oxidative decarboxylation of pyruvate with the help of the cofactor thiamine pyrophosphate (TPP). The other name for TPP, is vitamin B1: an essential nutrient. CO2 is also released in the first step of the PDH reaction(s). In the second reaction of PDH, catalysed by the E2 component, the two carbon product reacts with a molecular "swinging arm" comprising lipoic acid covalently coupled to a specific Lysine residue on the E2 enzyme, with a cyclic disulphide "head" group (see RHS). The acetylated lipoyl moiety is then exchanged with an incoming CoA molecule, thereby generating the major product of the reaction, acetyl CoA. The other product of this reaction is a reduced lipoic acid "swinging arm" which is re-oxidised by FAD; which is in turn converted to FADH. The latter reaction is catalysed by the third component of the complex, E3, or dihydrolipoamide dehydrogenase. This enzyme recycles FAD, through a coupled reduction of NAD to yield NADH. These three enzymatic steps make possible the transition from anaerobic metabolism to aerobic metabolism. The production of ATP in sufficient levels to drive the evolution of higher organisms, required a "quantum" increase in energy yield from the food that we and microbes digest: the PDH reaction is a key molecule in this evolutionary journey.
Let's now look at the structure-function relationships in PDH, which is the main focus of this particular post. You may have noticed that I mentioned a "swinging arm", and the three components E1, E2 and E3; all of which suggest something a little unusual. The first thing to draw your attention to is the size of PDH. There have been a number of Biochemists associated with the pioneering studies on the PDH reaction, but two names stand out: Lester Reed and Richard Perham (who sadly died earlier this year: you can read an appreciation here from Prof. Nigel Scrutton, or if you do not have full journal access here). Reed (who began publishing on the biochemistry of lipoic acid in the early 1950s, and Perham, a little later, devoted much of their early careers in purifying and characterising the PDH complex (a lovely perspective by Lester Reed is freely avalaible in J.Biol.Chem here). What became clear early on, was that this was a monster of an enzyme. To give you some perspective, the PDH complex (PDHc) is several times bigger than a ribosome.
Maths challenge. Calculate the volumes of E.coli (assuming a sphere of diameter 1.0μM, the volume of PDHc (assuming a sphere of diameter 0.03 x
10-3 μM) and the volume of a "regular" protein, such as haemoglobin tetramers (diameter 0.005 x 10-3μM).
Then divide the cell volume by the enzyme volume and estimate the
number of complexes that would fit inside the cell. Compare PDHc
with your haemoglobin calculations. For those who have forgotten, the
volume of a regular sphere can be calculated using the classical formula
4/3πr3
The unexpectedly large size of the PDHc meant on the one hand that X-ray diffraction would have been a significant challenge for an X-ray crystallographer in the 1960s. On the other hand, it was amenable to electron microscopy (albeit still in its infancy with biological macromolecules). In addition, the studies on the ribosome from other labs had made use of the then specialised method of analytical ultracentrifugation. This particular technique facilitated the otherwise difficult determination of oligomeric protein molecular weight by high speed sedimentation in a centrifuge that (at the time) was the size of a small car! Using these biophysical methods, Reed's group established that PDH was indeed a monster of a molecule. Some years later, the structures of sub-components of PDH began emerging (during the 1970s and '80s), and, partly as consequence of the introduction of recombinant DNA technology in the '80s and '90s, a combination of analysis of individual polypeptides and higher and higher resolution microscopy, the overall structural features of PDHc emerged (the eukaryotic PDHc is shown top left, from a lovely open access paper by Reed's group in Proc Nat Acad Sci).
The bacterial, and indeed yeast and human PDHcs comprise a large core of E2 molecules (recall this enzyme catalyses the transfer of the two carbon product of PDC to the lipoyl swinging arm). This assembly contains 24 or 60 copies of E2 ( RHS A is a 24mer and B a 60mer of the E2 core). The E1 heterotetramer
α2β2 dimer is present in multiple copies, with E3 being iterated 12 times. The overall complex has a subunit stoichiometry of 24:24:12 (E1:E2:E3) in bacteria and 30:60:12 in eukaryotes (and I am sure there will be species variation). In addition to these pioneering authors, Professor Michael Danson's
group at the University of Bath's Centre for Extremophile Research have spent years investigating structure
function relationships amongst several Krebs Cycle enzymes and the have often posed the question: why is
PDHc so big. See his laboratory's recent publication entitled "Why are the 2-oxo acid dehydrogenase complexes so large?"
 I
have mentioned before in a number of earlier posts, the power of
combining Molecular Genetics and Biochemistry. The unfolding of the PDHc
story provides another excellent example. In fact Professor John Guest
at Sheffield (who you can read about in his own words in an earlier post) set out to "clone" the Krebs Cycle genes over 30 years ago. This work, in parallel with that of Professor Richard Perham
in particular, unlocked the secrets of the "swinging arm" since the
primary structure motif is part of the substrate, and the number of arms
varies from organism to organism. The lipoyl
domain is attached directly to a conserved Lysine in the E2 core and it
migrates between the E1 and E3 active sites and its dynamic
characteristics were first investigated by Perham's group in collaboration with a number of UK NMR spectroscopists (notable Professor Gordon Roberts). It is important to note that even though teh PDHc is a "monster" and therefore not at all well suited to NMR analysis, highly dynamic subcomponents of large macromolecules can appear as distinctive features of the otherwise featureless NMR spectrum. The NMR structural ensemble for a bacterial lipoyl domain is shown (top LHS from Berg et al, 1996).
I
have mentioned before in a number of earlier posts, the power of
combining Molecular Genetics and Biochemistry. The unfolding of the PDHc
story provides another excellent example. In fact Professor John Guest
at Sheffield (who you can read about in his own words in an earlier post) set out to "clone" the Krebs Cycle genes over 30 years ago. This work, in parallel with that of Professor Richard Perham
in particular, unlocked the secrets of the "swinging arm" since the
primary structure motif is part of the substrate, and the number of arms
varies from organism to organism. The lipoyl
domain is attached directly to a conserved Lysine in the E2 core and it
migrates between the E1 and E3 active sites and its dynamic
characteristics were first investigated by Perham's group in collaboration with a number of UK NMR spectroscopists (notable Professor Gordon Roberts). It is important to note that even though teh PDHc is a "monster" and therefore not at all well suited to NMR analysis, highly dynamic subcomponents of large macromolecules can appear as distinctive features of the otherwise featureless NMR spectrum. The NMR structural ensemble for a bacterial lipoyl domain is shown (top LHS from Berg et al, 1996). The culmination of these investigations have revealed a fascinating picture of a "molecular machine" that delivers acetyl CoA to the active site of citrate synthase where it reacts with oxaloacetate to produce citrate. The citrate proceeds through a series of interconversions until it emerges again as oxaloacetate, ready to accept a second acetyl CoA molecule. The product of pyruvate decarboxylationis transferred to the lipoyl swinging arm to generate acetyl CoA. The lipoyl domain is then re-oxidised by FAD and the FADH produced is in turn re-oxidised by NAD, which is released as NADH and this goes on to generate ATP via the electron transfer chain present in the inner mitochondrial membrane (in eukaryotes).
The culmination of these investigations have revealed a fascinating picture of a "molecular machine" that delivers acetyl CoA to the active site of citrate synthase where it reacts with oxaloacetate to produce citrate. The citrate proceeds through a series of interconversions until it emerges again as oxaloacetate, ready to accept a second acetyl CoA molecule. The product of pyruvate decarboxylationis transferred to the lipoyl swinging arm to generate acetyl CoA. The lipoyl domain is then re-oxidised by FAD and the FADH produced is in turn re-oxidised by NAD, which is released as NADH and this goes on to generate ATP via the electron transfer chain present in the inner mitochondrial membrane (in eukaryotes).
There are some further features of the PDHc that I could discuss, in particular its regulation by phosphorylation. However, this would be for another post. I want to leave you with a thought regarding the lipoamide dehydrogenase (E3) component which is common to the the 2- oxoglutarate dehydrogenase complex, another multienzyme complex (which you will see discussed by Perham and Reed if you follow their work on Pubmed). In fact another two coli enzymes share the E3 enzyme. Think for a minute: one gene 4 enzymes, a slight variation on the "one gene: one enzyme hypothesis" associated with Archibald Garrod
and Beadle and Tatum! Now think about the evolutionary consequences of a
single enzyme, specifically articulating with four quite distinct
complexes. I think this is something for a conversation in the Innovation Labs next week?


